Prior Infections
A history of human infectious disease puts the probability of a COVID lab leak in perspective
A brand new human coronavirus emerging just half an hour’s drive from a lab that studies coronaviruses demands one consider the possibility that the lab could have been somehow responsible. At first glance, the chances that this was merely a coincidence seem vanishingly small. But a review of the history of novel pathogens and laboratory releases suggests that the chances of a lab accidentally introducing a new human pathogen are smaller still.
Specifically, the evidence will show that:
Every novel human pathogen ever identified has come from animals with the majority originating in wildlife.1
Laboratory releases are rare and have historically involved known pathogens that are in culture in the lab, not just samples collected by the lab.
The epidemiological signature of laboratory releases differs sharply from the pattern seen in COVID-19.
When a new disease is first identified, the pathogen has usually infected other people, often long before and far from the first official case.
Origin Stories
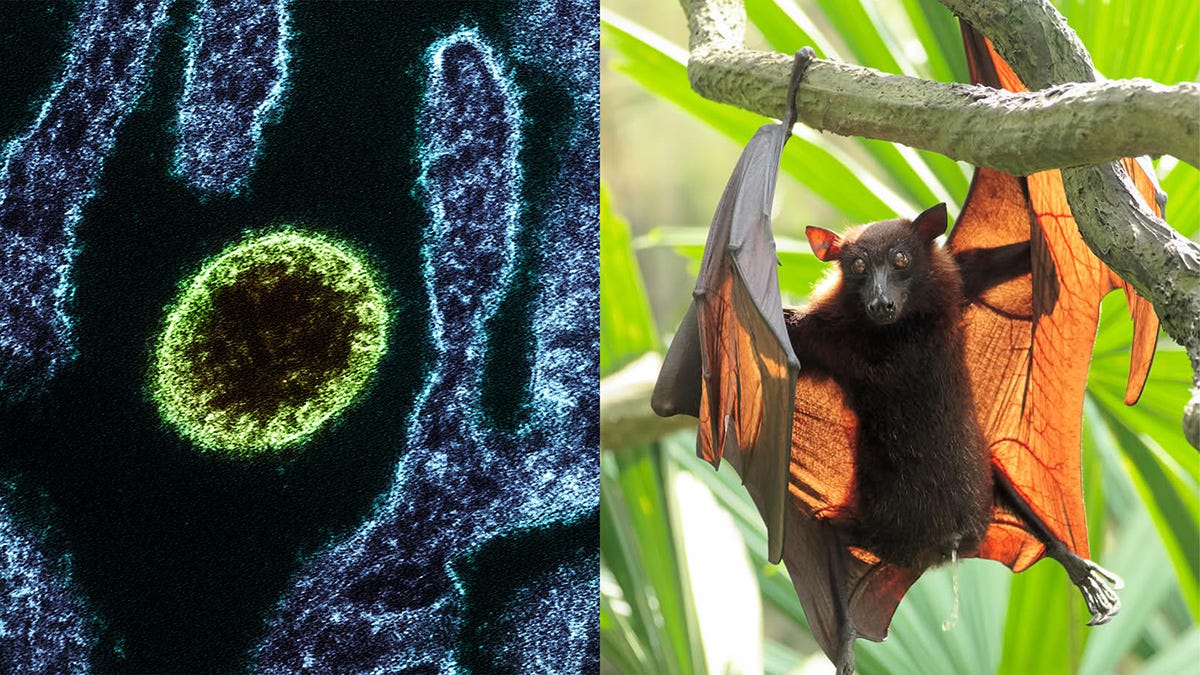
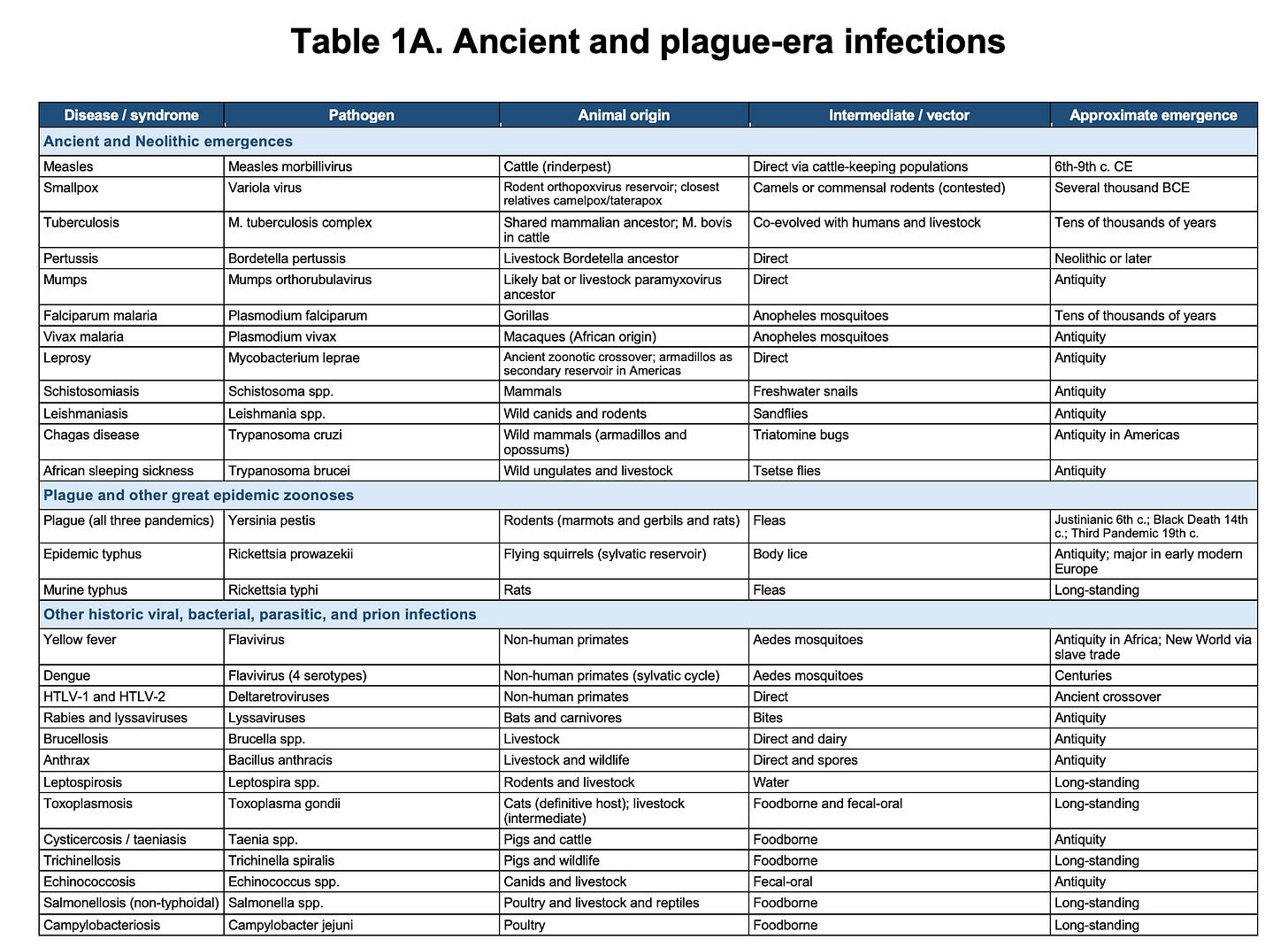
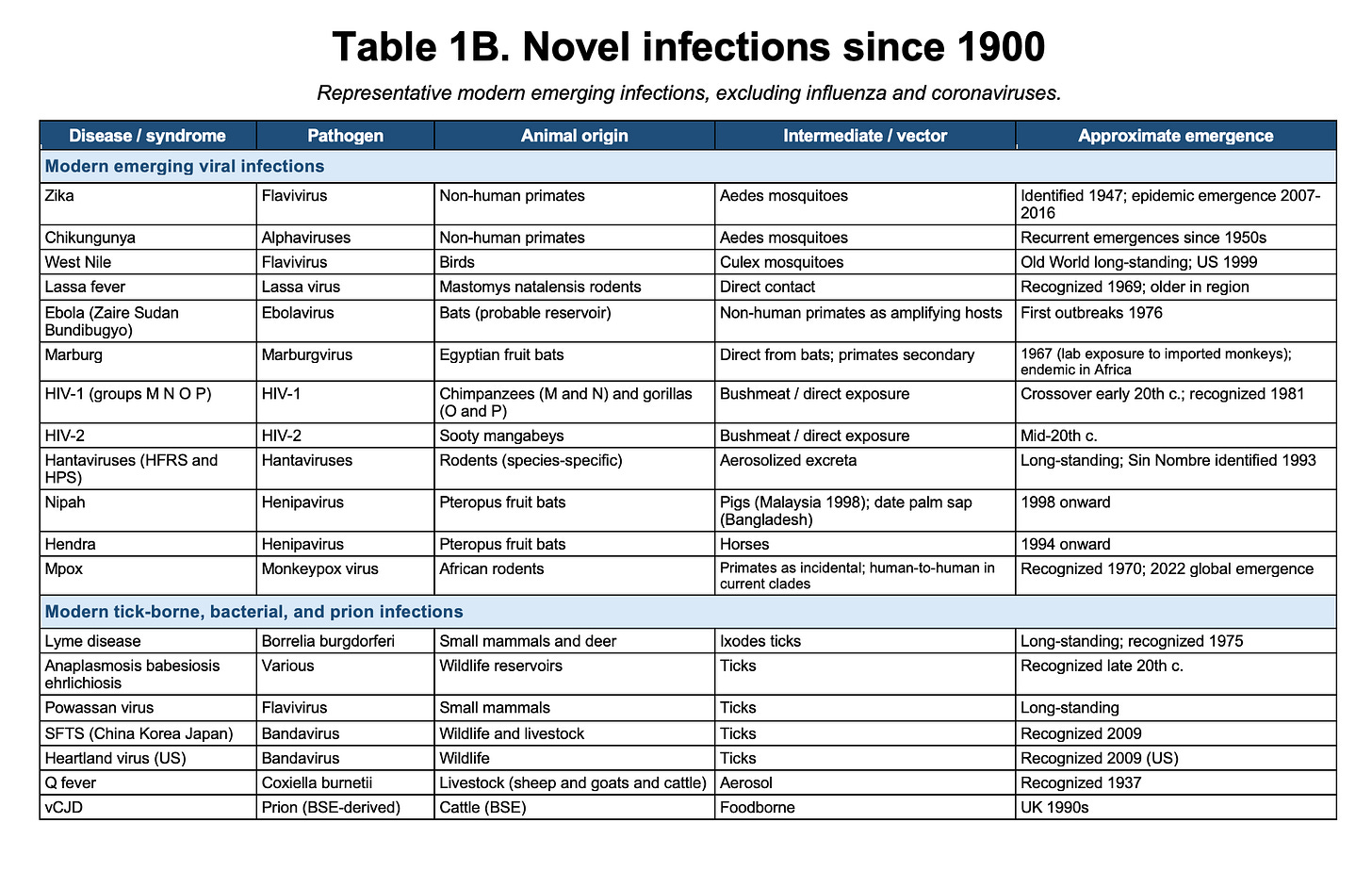
Every new human pathogen on record, without exception, has emerged from an animal reservoir. Measles evolved from a cattle virus. HIV emerged from chimpanzees and sooty mangabeys. The 1918, 1957, 1968, and 2009 influenza pandemics from avian and swine reassortment. SARS from bats through palm civets. MERS from bats through dromedary camels. Ebola, Marburg, Nipah, Hendra, hantaviruses, Lassa, mpox, yellow fever, dengue, Zika, chikungunya, plague, typhus, Lyme. The list runs from antiquity to last year’s H5N1 spillover into dairy cattle. It does not contain a single counterexample.
By “new human pathogen”, I mean a pathogen newly established in humans, not a known pathogen accidentally released from a laboratory or reintroduced after prior circulation. I have included a table with a representative historical cross-section of major human infectious diseases and emerging infections at the end of this post. Their importance lies not in any single example, but in the absence of counterexamples.
This is not a rhetorical flourish. It is the base rate. Every novel human infection on record has emerged from the natural environment. This pattern holds across millennia and across pathogen types: viruses, bacteria, parasites, and prions.
To argue that SARS-CoV-2 originated in a laboratory is to propose that a global pandemic arose from a lab leak for the first time in human history. Not the first undetected lab leak to cause a pandemic. The first lab leak of any kind to cause a pandemic. Ever. Laboratory infections and laboratory releases have occurred. But none, not one, has ever introduced a fundamentally new human pathogen.
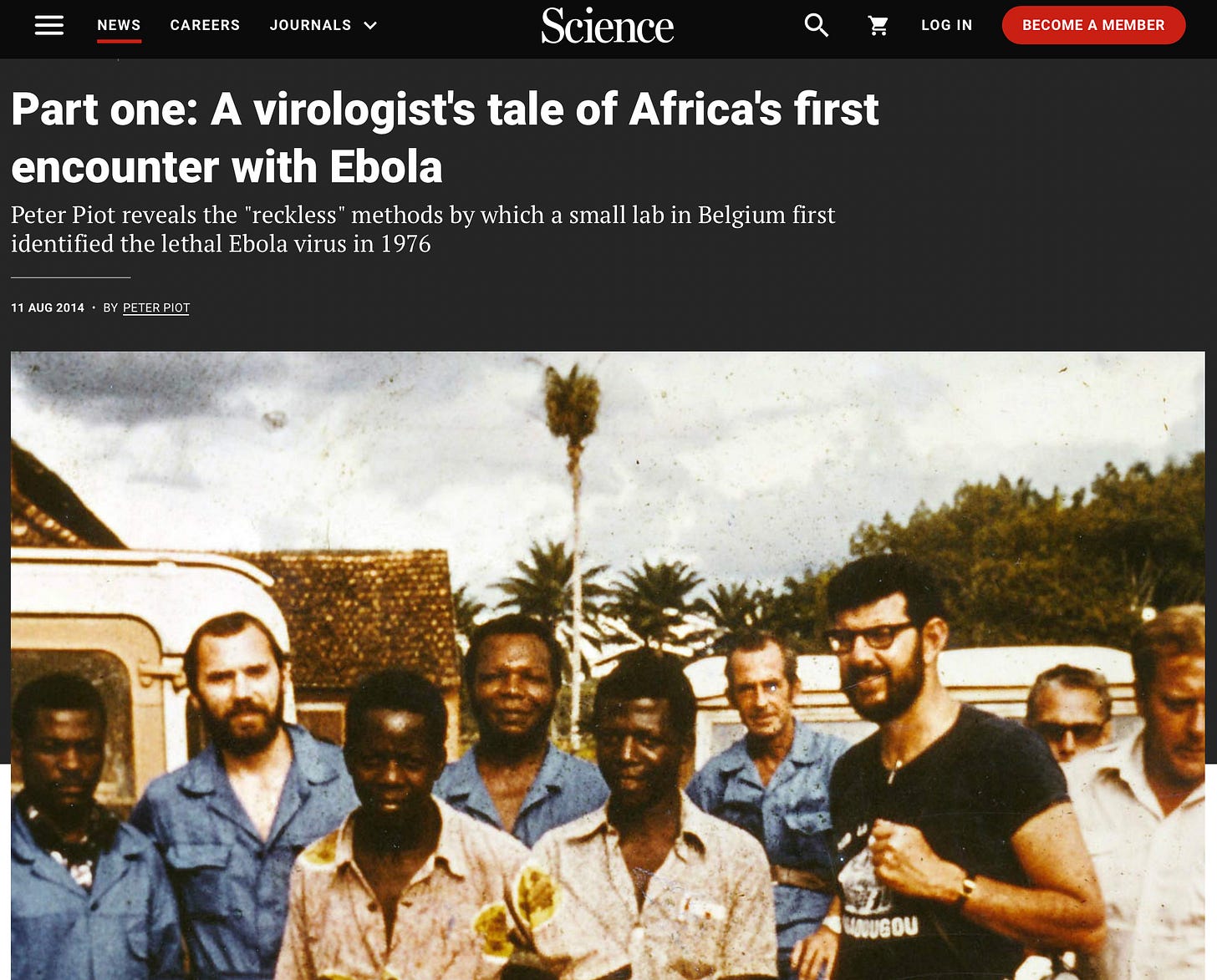
When a pathogen first jumps from an animal reservoir to humans, the epidemiological signature is unmistakable: initial cases cluster around the location where humans and animals come into contact. Plague in the 14th century emerged in Central Asia among hunters and herders with exposure to infected marmots and rodents, then spread along trade routes. When Ebola appeared in 1976, the first cases were in villages adjacent to rainforests in Sudan and Zaire, among people with contact to infected primates or bushmeat. When SARS-CoV-1 emerged in 2002, cases appeared in Guangdong Province among people with exposure to wildlife markets where live civets and raccoon dogs were sold, but the outbreak was not identified until it reached Hong Kong, a major international hub, in February 2003. When MERS began in 2012, the initial cases were in the Arabian Peninsula among people with direct contact to infected dromedary camels, particularly camel handlers and slaughterhouse workers. When H5N1 spilled over into United States dairy cattle in early 2024, the first infected farms were located on the Central North American migratory bird flyway, where wild birds and livestock come into direct contact.
In a zoonotic emergence, the origin is the spillover interface: the place where humans encounter the infected animal reservoir or intermediate host. What may differ is whether we identify that location immediately, reconstruct it later, or miss it entirely because the first infections occurred in a setting with little surveillance.
The spread pattern that follows is consistent: infections radiate outward from the spillover point through human-to-human transmission, animal movement, trade routes, or some combination of these. The epicenter of recognition may be a city, hospital, port, or market. But the biological origin remains the place where humans first encountered the infected animal source.
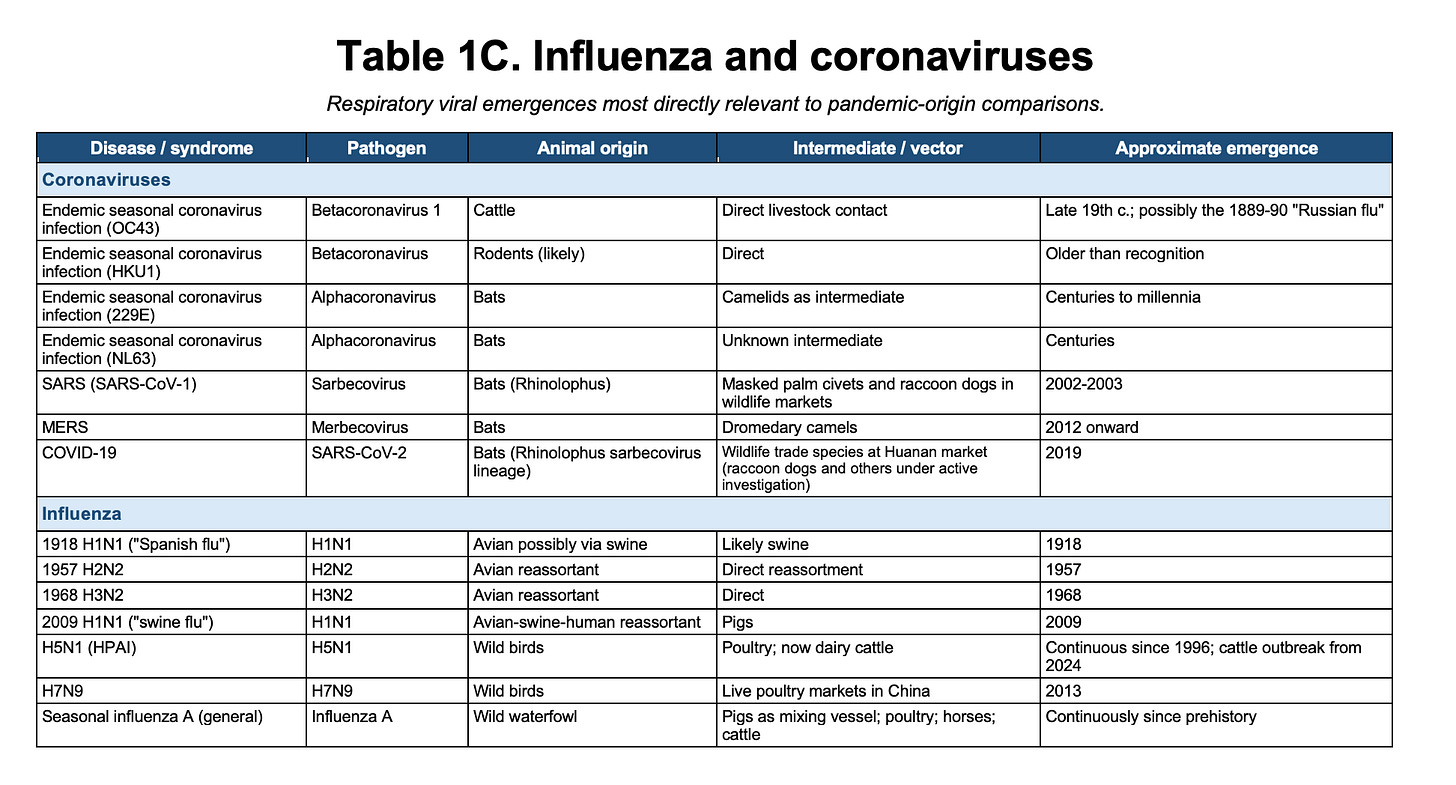
Coronaviruses Are Not an Exception
Coronaviruses fit this same pattern. They are not a unique class of pathogens requiring a different historical framework. The endemic seasonal human coronaviruses, including OC43, HKU1, 229E, and NL63, all point back to animal reservoirs, possibly bats. They are often called “common cold” coronaviruses because that is what they usually cause today, but that does not mean they were mild when they first entered humans. OC43, for example, is closely related to bovine coronaviruses and may have crossed into humans in the late nineteenth century. Some have suggested a possible connection to the 1889–1890 “Russian flu” pandemic, though that remains unresolved.
The severe human coronaviruses follow the same pattern. SARS-CoV-1 emerged from a bat coronavirus lineage through wildlife-market animals. MERS-CoV is maintained in dromedary camels, with bats likely representing the deeper evolutionary reservoir. The details differ, and the precise intermediate pathway is not always known, but the pattern is familiar: animal reservoir, ecological contact, spillover, and, in rare cases, sustained human transmission.
Coronaviruses therefore reinforce the broader historical pattern rather than challenging it. They are not evidence that COVID-19 belongs in a special category. They are evidence that the emergence of a novel human coronavirus should be evaluated against the same zoonotic history as every other novel human infection.
What Laboratory Releases Actually Look Like
There are documented cases in which pathogens escaped from laboratories and infected people in the community. They tell a coherent story about what actual lab leaks look like and how the research community has responded to them. There are two plausible ways for a pathogen to spread from a lab to the general population: direct environmental release or infection of a lab worker (or escape of an infected lab animal) followed by human-to-human transmission. Each has a signature.
Large containment breaches with environmental spread
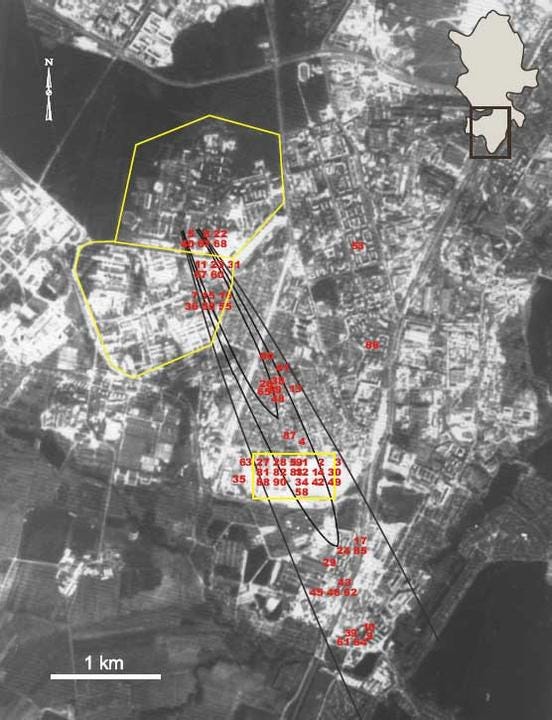
In 1979, a Soviet military microbiology facility in Sverdlovsk, the Institute of Ultra-Pure Biological Preparations, had a containment failure in its air handling system. Aerosolized Bacillus anthracis escaped into the environment. The outbreak killed an estimated 60 to over 100 people, though exact numbers remain classified in Russian records. It was initially covered up by Soviet authorities, but investigations after the Cold War established the lab origin definitively.
More recently, in 2019, inadequate inactivation procedures at a vaccine manufacturing facility in Lanzhou, China, caused it to release aerosolized Brucella melitensis through its exhaust system. Over 3,000 people in surrounding communities were infected. The outbreak was identified through epidemiological investigation, traced to the facility, and contained through public health measures.
In a non-human example, in 2007 a lab the UK that was studying Foot and Mouth disease, accidentally released contaminated wastewater, causing infections in four nearby farms.
All of these outbreaks were defined by a geographically concentrated plume of illness in the immediate vicinity of the facility. The source was not a mystery epidemiologically, even when authorities tried to obscure it.
Infected lab worker outbreaks: rapid identification and containment
The second pattern involves lab workers becoming infected and transmitting to contacts. This happened with SARS-CoV-1 on three documented occasions after the 2003 outbreak was controlled: Singapore in September 2003, Taiwan in December 2003, and the Beijing National Institute of Virology in April 2004.
Critically, none of these became large. Each was identified within days or weeks of the first case because the infected person was a researcher working with the virus, they recognized their own symptoms, sought medical attention, and the hospital immediately flagged it as a known pathogen. Contact tracing was swift. The largest outbreak, Beijing, involved nine secondary cases total. All were contained before community transmission could establish.
This is the consistent pattern in the documented SARS laboratory infections, and it is what one would expect when a trained researcher becomes infected with a known high-consequence pathogen. The outbreak cannot grow large because identification happens in real time. Few people are more likely to notice a suspicious respiratory illness than someone working with potential respiratory pathogens.
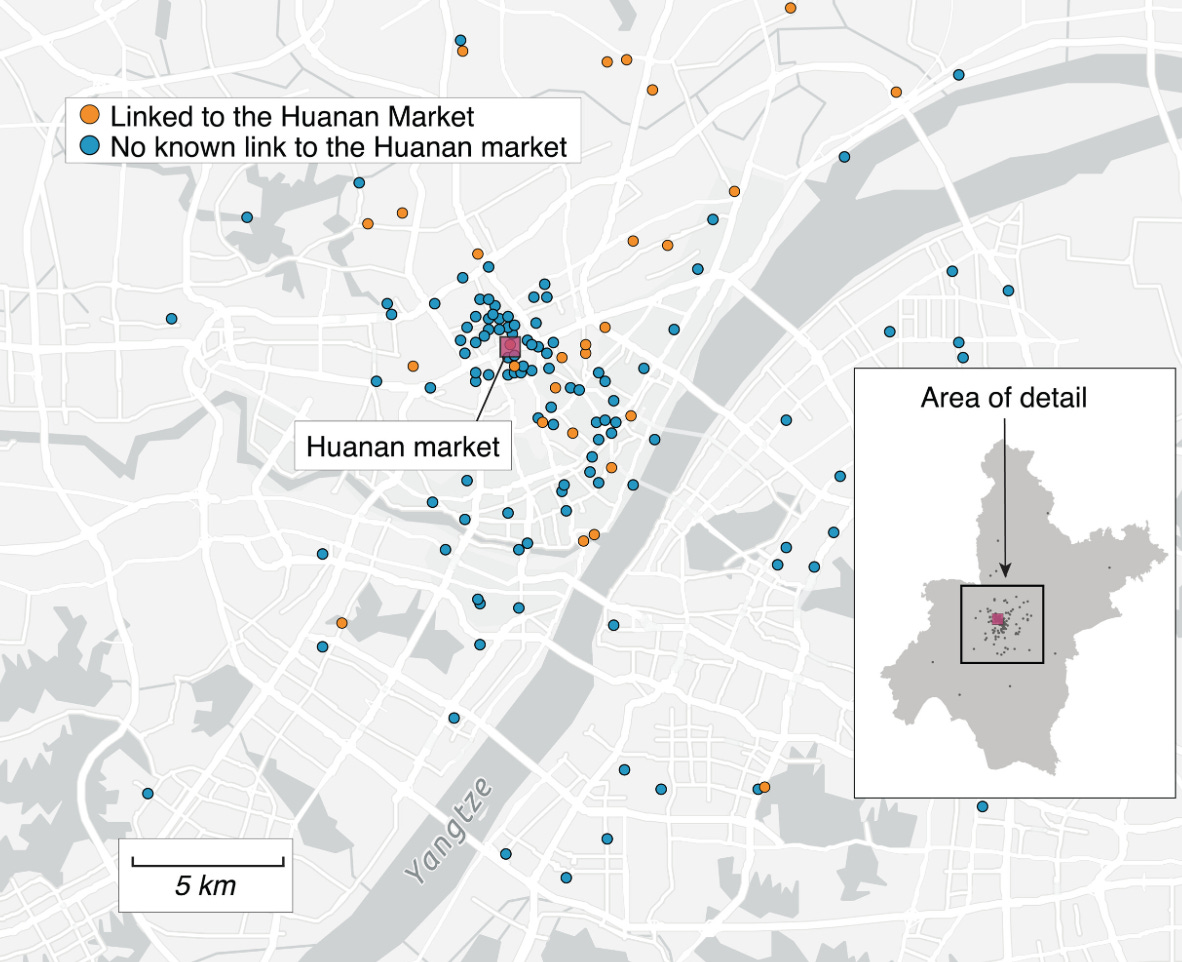
Of two possible models for laboratory origin, COVID-19 fits neither. No laboratory worker presented to a hospital with a novel respiratory illness in December 2019 that was recognized as SARS-like and triggered immediate investigation. There were no clusters identified around the homes of laboratory workers. The earliest known cases clustered around a wildlife market selling live mammals from species groups known to be susceptible to SARS-related coronaviruses. It spread for weeks before anyone identified it as novel. By the time the world’s coronavirus researchers were looking at it, it had already seeded a pandemic.
Why Wuhan?
Proponents of a lab leak argue that the appearance of a novel coronavirus in Wuhan within a few miles of a lab studying novel coronaviruses is too great a coincidence. This is a legitimate question that merits careful consideration. Again, let’s put it in the context of known zoonoses.
In considering the epidemiology of a novel pathogen, it is critical to differentiate between the first case and the first identified case. In all likelihood, they will not be the same. Recognition of a human infection from a novel pathogen depends on the interplay among five factors: severity, transmissibility, rapidity of symptom onset, population density, and availability of diagnostic capacity to identify novel pathogens. Reducing any one of these can obscure the pathogen. SARS-CoV-1 is instructive here. Cases appeared in Guangdong Province in late 2002 among people with exposure to wildlife markets, but the outbreak went unrecognized for weeks. The virus was spreading at the spillover interface. We simply could not see it until it reached Hong Kong, a major international hub with sophisticated medical surveillance, and then spread to other cities. The first identification occurred in a population center, not at the origin.
Ebola jumped to humans in remote areas of Sudan and Congo in 1976 and was immediately recognized because it had high severity and relatively rapid symptom onset. People died within days. This dramatic acute illness got noticed even in villages with minimal medical infrastructure. High severity compensated for low population density and limited diagnostic capacity.
HIV crossed from chimpanzees to humans in Central Africa probably in the early 20th century through bushmeat hunting. It had low initial severity, slow disease progression, and low transmissibility in the early stages. People got gradually sick over years, developed opportunistic infections, and died, but individually these looked like routine illnesses in dispersed populations. The virus remained unidentified until it reached dense urban populations in New York and San Francisco in the 1980s, where high population density and better diagnostic infrastructure created a recognizable epidemic pattern. The virus had been spreading through Central Africa for 60 years. We simply could not see it.
COVID-19 is highly transmissible with relatively rapid symptom onset, but moderate severity. In a rural area with low population density and limited diagnostic capacity, a person gets sick with pneumonia that looks like seasonal flu, recovers or dies, and no one recognizes it as something novel. The moderate severity means it blends into background respiratory illness. COVID-19’s high transmissibility could create clusters even in rural areas, yet without population density and diagnostic capacity, those clusters still go unrecognized as novel. Only when COVID-19 reached Wuhan did all five factors align for detection. Wuhan was a major urban center with high population density, a crowded wildlife market where live animals and human handlers were in close contact, and a world-class virology laboratory uniquely well-equipped to identify a novel coronavirus.
The pattern defined by other zoonotic infections suggests that the first human cases of COVID-19 may well have occurred far from Wuhan, weeks or months earlier, in rural areas where people had contact with infected animals. Those cases may have gone unrecognized because moderate severity made the disease indistinguishable from routine illness, rural areas lack population density for rapid spread, and the capacity to identify a novel pathogen. Alternatively, it was transported there in the animals bred for the markets in rural farms. Wuhan had a perfect setting for explosive spread in the Huanan market and one of the best labs in the world for identifying a novel coronavirus.
What the Pattern Tells Us
The history of human infectious disease is a history of contact with animals. Domestication brought us measles and smallpox and tuberculosis. The slave trade carried yellow fever across an ocean. Bushmeat hunting gave us HIV. Deforestation, forest fragmentation, and wildlife trade have repeatedly opened new pathways for spillover: Ebola from disrupted forest ecologies, Nipah from fruit bats into pigs and people, SARS through the wildlife trade, mpox through wildlife contact and animal movement, and zoonotic malaria where forest loss brings humans, macaques, and mosquitoes into new alignment. Microbes are tightly woven into the fabric of wild ecosystems. Human disruption of those ecosystems inevitably exposes us to new potential pathogens.
The point is not to decide the origin of COVID-19 by assertion. It is to define the historical universe into which any proposed origin must fit. Across that universe, novel human pathogens have emerged from animal reservoirs. Laboratory releases, when they have occurred, have involved known pathogens and have followed recognizable patterns.
COVID-19 should be evaluated against this background. A laboratory origin would not merely be another example of a familiar category. It would require two categories of events not previously documented: the laboratory introduction of a fundamentally new human pathogen and a laboratory release of any pathogen causing a global pandemic.
In other words, the lab-leak hypothesis, as configured by its proponents, must overcome a historical record with no known precedent. That requires extraordinary evidence.
The burden of proof has always belonged to the people making the extraordinary claim. In the case of COVID-19, that burden has not been met.
References
For the great majority of the diseases referenced above, the wildlife or livestock origin is established in standard infectious disease references such as Bennett JE, Dolin R, Blaser MJ, eds. Mandell, Douglas, and Bennett’s Principles and Practice of Infectious Diseases. 9th ed. Philadelphia: Elsevier; 2020; and the CDC’s National Center for Emerging and Zoonotic Infectious Diseases (https://www.cdc.gov/ncezid). The references below are limited to claims that are recently established, contested, or otherwise warrant primary citation.
Furuse Y, Suzuki A, Oshitani H. Origin of measles virus: divergence from rinderpest virus between the 11th and 12th centuries. Virol J. 2010;7:52. doi:10.1186/1743-422X-7-52
Mühlemann B, Vinner L, Margaryan A, et al. Diverse variola virus (smallpox) strains were widespread in northern Europe in the Viking Age. Science. 2020;369(6502):eaaw8977. doi:10.1126/science.aaw8977
Li Y, Carroll DS, Gardner SN, Walsh MC, Vitalis EA, Damon IK. On the origin of smallpox: correlating variola phylogenics with historical smallpox records. Proc Natl Acad Sci USA. 2007;104(40):15787-15792. doi:10.1073/pnas.0609268104
Taubenberger JK, Morens DM. 1918 influenza: the mother of all pandemics. Emerg Infect Dis. 2006;12(1):15-22. doi:10.3201/eid1201.050979
Worobey M, Han GZ, Rambaut A. A synchronized global sweep of the internal genes of modern avian influenza virus. Nature. 2014;508(7495):254-257. doi:10.1038/nature13016
Caserta LC, Frye EA, Butt SL, et al. Spillover of highly pathogenic avian influenza H5N1 virus to dairy cattle. Nature. 2024;634(8034):669-676. doi:10.1038/s41586-024-07849-4
Vijgen L, Keyaerts E, Moës E, et al. Complete genomic sequence of human coronavirus OC43: molecular clock analysis suggests a relatively recent zoonotic coronavirus transmission event. J Virol. 2005;79(3):1595-1604. doi:10.1128/JVI.79.3.1595-1604.2005
Guan Y, Zheng BJ, He YQ, et al. Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science. 2003;302(5643):276-278. doi:10.1126/science.1087139
Li W, Shi Z, Yu M, et al. Bats are natural reservoirs of SARS-like coronaviruses. Science. 2005;310(5748):676-679. doi:10.1126/science.1118391
Azhar EI, El-Kafrawy SA, Farraj SA, et al. Evidence for camel-to-human transmission of MERS coronavirus. N Engl J Med. 2014;370(26):2499-2505. doi:10.1056/NEJMoa1401505
Mohd HA, Al-Tawfiq JA, Memish ZA. Middle East Respiratory Syndrome Coronavirus (MERS-CoV) origin and animal reservoir. Virol J. 2016;13:87. doi:10.1186/s12985-016-0544-0
Worobey M, Levy JI, Malpica Serrano L, et al. The Huanan Seafood Wholesale Market in Wuhan was the early epicenter of the COVID-19 pandemic. Science. 2022;377(6609):951-959. doi:10.1126/science.abp8715
Pekar JE, Magee A, Parker E, et al. The molecular epidemiology of multiple zoonotic origins of SARS-CoV-2.Science. 2022;377(6609):960-966. doi:10.1126/science.abp8337
Holmes EC, Goldstein SA, Rasmussen AL, et al. The origins of SARS-CoV-2: a critical review. Cell. 2021;184(19):4848-4856. doi:10.1016/j.cell.2021.08.017
Sharp PM, Hahn BH. Origins of HIV and the AIDS pandemic. Cold Spring Harb Perspect Med. 2011;1(1):a006841. doi:10.1101/cshperspect.a006841
Keele BF, Van Heuverswyn F, Li Y, et al. Chimpanzee reservoirs of pandemic and nonpandemic HIV-1. Science. 2006;313(5786):523-526. doi:10.1126/science.1126531
World Health Organization. Mpox. WHO Fact Sheet. Updated 2024. https://www.who.int/news-room/fact-sheets/detail/mpox
On the 1977 H1N1 lab origin and SARS-CoV-1 lab escapes
Rozo M, Gronvall GK. The reemergent 1977 H1N1 strain and the gain-of-function debate. mBio. 2015;6(4):e01013-15. doi:10.1128/mBio.01013-15
Wertheim JO. The re-emergence of H1N1 influenza virus in 1977: a cautionary tale for estimating divergence times using biologically unrealistic sampling dates. PLoS One. 2010;5(6):e11184. doi:10.1371/journal.pone.0011184
Kalyar F, Chughtai AA, MacIntyre CR. Origin of the H1N1 (Russian influenza) pandemic of 1977—a risk assessment using the modified Grunow-Finke tool. Risk Anal. 2024;44(11):2484-2494. doi:10.1111/risa.14343
World Health Organization. China’s latest SARS outbreak has been contained, but biosafety concerns remain. Press release. 18 May 2004. https://www.who.int/news/item/18-05-2004-china-s-latest-sars-outbreak-has-been-contained-but-biosafety-concerns-remain
Lim PL, Kurup A, Gopalakrishna G, et al. Laboratory-acquired severe acute respiratory syndrome. N Engl J Med. 2004;350(17):1740-1745. doi:10.1056/NEJMoa032565
Normile D. Mounting lab accidents raise SARS fears. Science. 2004;304(5671):659-661. doi:10.1126/science.304.5671.659
G.T. Keusch, J.H. Amuasi, D.E. Anderson, P. Daszak, I. Eckerle, H. Field, M. Koopmans, S.K. Lam, C.G. Das Neves, M. Peiris, S. Perlman, S. Wacharapluesadee, S. Yadana, & L. Saif, Pandemic origins and a One Health approach to preparedness and prevention: Solutions based on SARS-CoV-2 and other RNA viruses, Proc. Natl. Acad. Sci. U.S.A. 119 (42) e2202871119, https://doi.org/10.1073/pnas.2202871119 (2022).
On documented containment breaches: anthrax and Brucella
Guillemin J. American Anthrax: Fear, Crime, and the Investigation of the 2001 Bioterrorism Attacks. New York: Metropolitan Books; 2011. [Detailed historical and investigative account of the 1979 Sverdlovsk release and subsequent cover-up]
Meselson M, Guillemin J, Hugh-Jones M, et al. The Sverdlovsk anthrax outbreak of 1979. Science. 1994;266(5188):1202-1208. doi:10.1126/science.7973702
Cui B, Smajlovic S, Ernst M, et al. Isolation and characterization of Brucella melitensis from the 2019 Lanzhou vaccine facility incident. Emerg Infect Dis. 2020;26(12):2877-2880. doi:10.3201/eid2612.201433
Pappas G, Akritidis N, Christou L. Brucellosis. Lancet. 2006;367(9516):1139-1142. doi:10.1016/S0140-6736(06)68433-4
On vCJD
Will RG, Ironside JW, Zeidler M, et al. A new variant of Creutzfeldt-Jakob disease in the UK. Lancet. 1996;347(9006):921-925. doi:10.1016/s0140-6736(96)91412-9
Bruce ME, Will RG, Ironside JW, et al. Transmissions to mice indicate that ‘new variant’ CJD is caused by the BSE agent. Nature. 1997;389(5650):498-501. doi:10.1038/39057
Note that I have not included fungal infections, which come from the natural environment, generally not from animals.


What an excellent, succinct article that lays out the logic of the so-called 'zoonotic origin' theory, and the clear implausibility of the so-called 'lab leak' theory. We're currently in the midst of another zoonotic disease outbreak caused by Andes hantavirus, and despite the obvious pathway for exposure (birdwatchers having visited the countries where the wildlife reservoirs exist), already some are proposing it emerged from a lab. In fact, many recent zoonotic disease emergence events have led to allegations of lab leak (e.g. Ebola) or vaccine origin (e.g. HIV). It seems that these conspiracy theories strike a chord with some people and are put forward to help them cope with the seeming serendipity of nature.
Robert, do you (through Eco Logic) have any current or former clients who have requested you to write on this topic?